Next generation sequencing presents new challenges in large scale data processing. In collaboration with the University of Liverpool’s Animal Sciences & Physiology Research group, in particular Dr Harry Noyes, we combined Taverna scientific workflows with computing power from the Amazon cloud to create a powerful next generation sequencing application for whole genome Single Nucleotide Polymorphism (SNP) analysis.
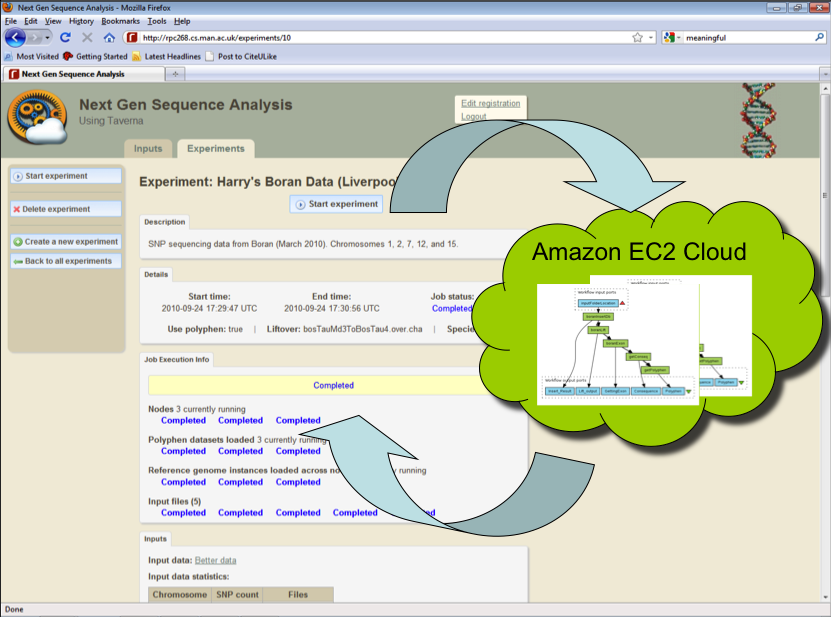
Through a Web portal, the application allows scientists to upload their input data, fire off a number of parallel cloud instances for the analysis, monitor progress and collect results (see figure below).
Preliminary work on the genetic variation of African cattle showed we can run a whole genome of ~22 million SNPs in a matter of hours. This work focuses on the response to trypanomiasis infection (sleeping sickness) in different cattle species.
The application was demonstrated at the European Conference of Computational Biology (ECCB) 2010, Ghent, Belgium, under the title “Software for the Data-Driven Researcher of the Future” – see the slides and video (no audio or subtitles yet!) from the talk.
This cloud application was based on the next generation sequencing work done presented at Bioinformatics Open Source Conference (BOSC) 2010, Boston, USA, under the title “Analysing African and European cattle with Taverna 2.2″. See the slides from the talk.